

Wet-lab profiling and computational informatics: two inseparable halves of our breast cancer program
The Hallett Lab has a long-standing program in breast cancer that spans both the genomics — the generation and processing of molecular data from clinical cohorts — and the informatics — the development of computational methods to extract insights from that data. These two activities are inseparable: the challenges of profiling real patient material directly motivate new analytic tools, and those tools open new biological questions requiring better profiling.
Our goals are both clinically actionable findings that guide treatment decisions and a fundamental understanding of breast cancer biology. This work is tightly integrated with the Dumeaux lab and with clinical partners across Canada, Norway, Sweden and the United States.
{/* Subtheme 1 */}Breast cancer is not one disease. Intrinsic subtypes — Luminal A, Luminal B, HER2-enriched, Basal-like, and Normal-like — reflect distinct biological programs with markedly different clinical behaviours. Our AIMS and AIPS packages assign patients to these subtypes at the level of individual samples, without requiring cohort-level normalization.
Subtype alone does not tell the full story. In collaboration with the Park lab at McGill, we showed that gene expression in the tumor stroma is strongly predictive of clinical outcome. Our STROMA4 package identifies four distinct stromal subtypes — T-cell infiltration, B-cell infiltration, stromal epithelial infiltration, and desmoplasia — adding significant prognostic information beyond tumor subtype.
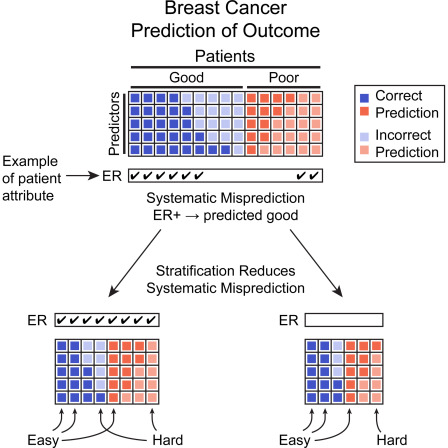
A recurring insight is that the tumor alone is not sufficient to understand clinical outcome. We showed that patients stratify into groups where outcome is easy to predict from molecular features, and groups where it is inherently difficult — reflecting genuine biological complexity. This introduced the concept of inherent prognostic difficulty in breast cancer.
Going beyond the tumor, we showed that gene expression in peripheral blood carries independent and complementary prognostic information to the primary tumor. The MIxT framework — developed in collaboration with Vanessa Dumeaux — enables principled comparison of transcriptional programs across matched tissues. Interactive explorations of these results are available here:
Ductal carcinoma in situ (DCIS) is a non-obligate precursor to invasive breast cancer. We lack robust biomarkers to distinguish indolent from progressive DCIS at diagnosis. A powerful example of our wet-to-dry feedback is PREFFECT: experience profiling archival FFPE tissue revealed how severely formalin fixation degrades RNA, motivating us to build a deep generative model that imputes high-quality transcriptomic profiles from FFPE material.
High-throughput CRISPR perturbation with single-cell readout to infer causal biological networks
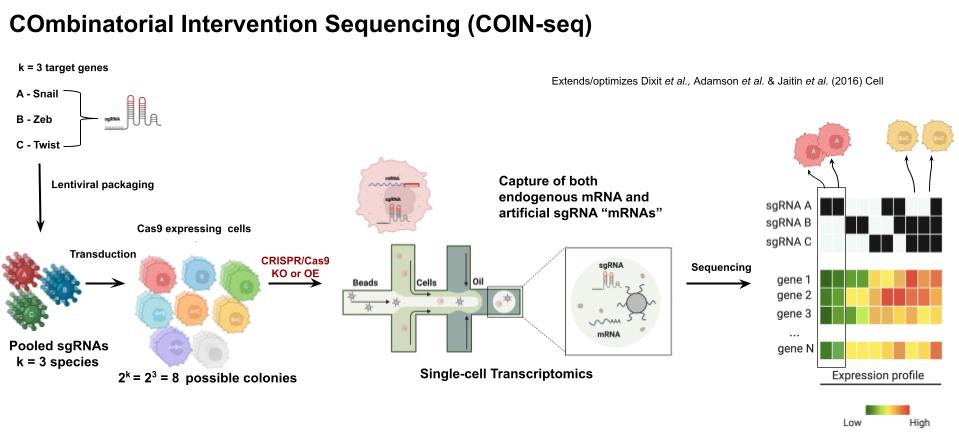
Understanding the causal relationships between genes is one of the central problems in molecular biology. Our COIN-seq platform delivers combinations of CRISPR/Cas9-mediated perturbations via lentiviral vectors and reads out the consequences at single-cell resolution. Each cell receives a pseudorandom subset of the perturbation library, allowing probabilistic and deep learning models to infer the underlying gene regulatory network at scale.
Single-cell profiling and deep-learning morphology analysis for a human fungal pathogen
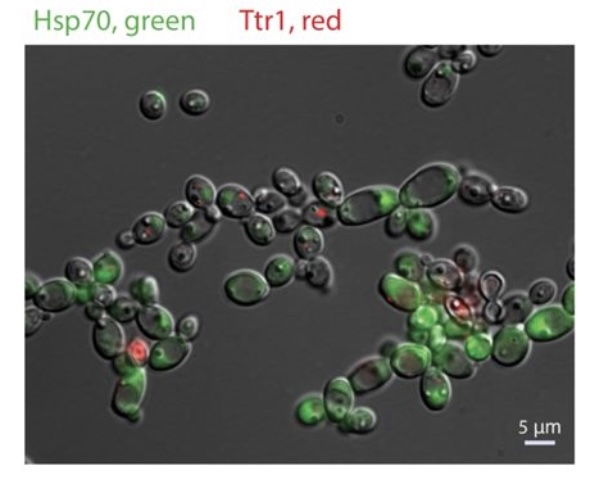
Candida albicans is the most common human fungal pathogen. Our research spans two interconnected directions: deep learning tools to quantify fungal morphology (Candescence), and single-cell profiling to understand phenotypic heterogeneity in drug-treated populations.
Candescence is built on a multi-task convolutional neural network (CNN) that jointly performs instance segmentation and morphological classification of C. albicans cells. Trained on annotated yeast, pseudohyphal, and hyphal forms, it uses focal loss, data augmentation, and a two-stage detect-then-classify pipeline to handle the extreme class imbalance and morphological continuum of fungal populations.
We adapted DROP-seq for C. albicans to profile transcriptional heterogeneity at single-cell resolution. In a large collaborative study in eLife, we identified distinct subpopulations with heterogeneous and adaptive cytoprotective responses to antifungal compounds — including stress response programs, efflux pumps, and metabolic reprogramming — providing a high-resolution map of drug tolerance diversity.
Generative models, neural architectures, and probabilistic methods applied across our research programs
Deep learning and generative modelling are not peripheral to our work — they are central to how we analyze molecular data across all four of our research programs. Probabilistic modelling, Bayesian inference, generative modelling and neural network theory are core to our research goals.
{/* Candescence */}
Candescence is built on a multi-task convolutional neural network (CNN) architecture that jointly performs instance segmentation and morphological classification of Candida albicans cells from microscopy images. The model is trained on a curated dataset of annotated yeast, pseudohyphal, and hyphal forms using a combination of pixel-wise segmentation loss and multi-class classification objectives. A key methodological challenge was handling the extreme class imbalance and morphological continuum in fungal populations — addressed through data augmentation, focal loss, and a two-stage detect-then-classify pipeline. The model produces per-cell morphological scores, organelle segmentation masks, and population-level summary statistics that enable rigorous statistical comparison across experimental conditions.
PREFFECT (Probabilistic Reconstruction of Expression From FFPE using Encoder-decoder Convolutional Transformers) frames FFPE transcriptomics correction as a conditional generative modelling problem. The core architecture is a variational autoencoder (VAE) in which the encoder maps degraded FFPE expression profiles into a structured latent space, and the decoder reconstructs a clean, fresh-frozen-equivalent profile. Crucially, PREFFECT incorporates an iterative refinement loop: the decoded output is re-encoded and refined over multiple passes, progressively removing FFPE-specific artifacts while preserving biologically meaningful variation. The model is trained with a combination of reconstruction loss, KL divergence regularization, and a contrastive loss that encourages matched FFPE/fresh-frozen pairs from the same tumor to cluster in latent space. This allows the model to disentangle technical degradation from true biological signal.
Deep learning is central to the analysis of COIN-seq data. After single-cell sequencing of combinatorially perturbed cells, we must infer which perturbations caused which transcriptional changes — and ultimately reconstruct the causal gene regulatory network. This requires models that can handle high-dimensional, sparse, and combinatorially structured data. We use graph neural networks and probabilistic deep learning to learn transcriptional signatures of individual perturbations, predict the effects of unseen combinations, and infer network structure from the ensemble of observed responses.
Beyond these specific applications, the lab maintains a broad interest in probabilistic modelling, Bayesian inference, generative modelling, and neural network theory — including variational autoencoders, normalizing flows, and generative models for single-cell data. We also maintain an active interest in agentic AI systems for accelerating scientific workflows. Our generic_cc infrastructure is a shared Claude Code environment designed for the lab's day-to-day use.